The role of ‘omics has almost completely shifted microbiologists approach to studying microbiomes. Prior to the introduction of ‘omics analysis techniques, microbiologists would commonly only be able to implement pure culture and chemical analysis techniques. ‘Omics analyses can be applied through techniques such as PCR and FISH.
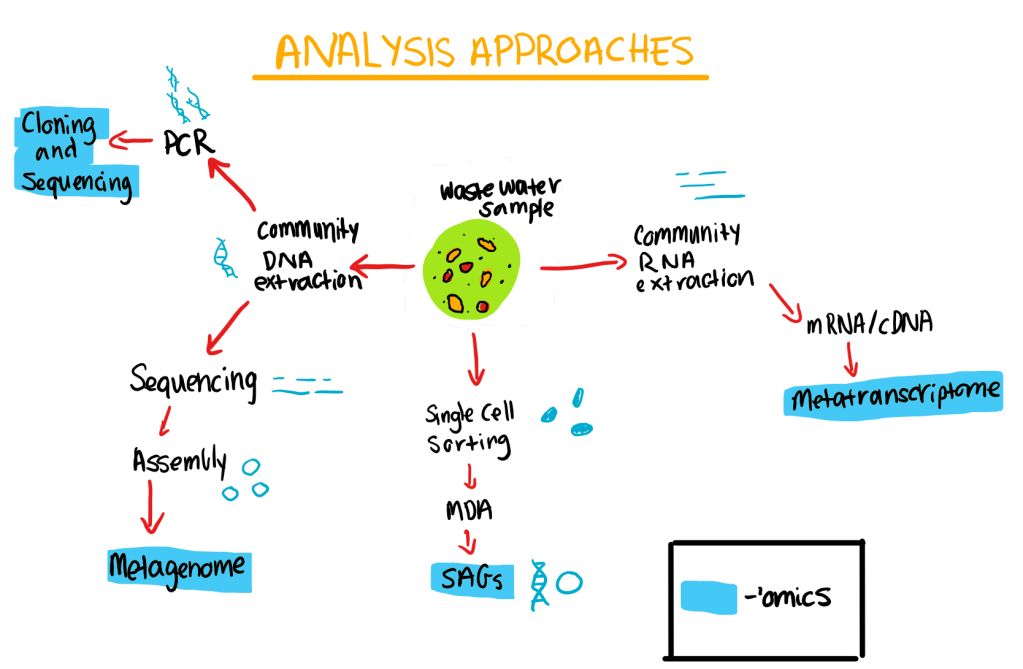
According to Ferrera and Sanchez (2016), the Polymerase Chain Reaction (PCR), had completely revolutionised the study of microorganisms as it allowed the analysis and identification of genes in non-pure culturable organisms. Often PCR can be used to identify microorganisms involved in key processes that occur in wastewater treatment plants such as carbon degradation, ammonia oxidation and reduction of nitrate. However, PCR can be limited by primer design for diverse metabolic or phenotypic genes being challenging (Ferrera and Sanchez, 2016).
Metagenomic studies can be useful to determine what microorganisms are present in a sample from a microbiome. Within the metagenome, it is possible to search for a functional gene. For example, it is possible to find genes involved in nitrogen removal such as nifH. Identifying the presence of nifH in a species or genus can suggest that the microorganism is involved in the nitrogen removal process, but these findings can potentially supported by an analysis of the metatranscriptome under the right environmental conditions. An example of such is a study conducted by Stokholm-Bjerregaard et al. (2017) where the genomes of microorganisms in 18 full-scale wastewater treatment plants were sequenced. It was found that an uncultured cyanobacteria by the name of “Candidatus Oscuribacter phosphatis” had the genes for the low-affinity inorganic phosphate transporter. The presence of this gene, however, was not sufficient evidence to conclude that this organism was a polyphosphate accumulating organism (PAO) involved in the Enhanced Biological Phosphorus Removal (EBPR) process. This was reinforced by the fact that other non-PAO organisms also had this sequence in their genome (Stokholm-Bjerregaard et al., 2017).
A metatranscriptome is a collection of all the genes in a sample that are transcripted (Fererra and Sanchez, 2016). This analysis is specific to the sample’s environment so it is possible to identify the microorganisms that produce proteins useful to wastewater treatment processes.
Ferrera and Sanchez (2016) recommend greater investment into sequencing and a combination of metagenomics and metatranscriptomics to provide better analysis of gene expression patterns in wastewater treatment systems. Although, it is agreed that metagenomic and metatranscriptome analysis approaches are limited due to biased information from the most abundant species in the sample. But using these techniques is also useful for circumventing isolation and PCR (Fererra and Sanchez, 2016).
‘Omics in microbiology are designed to work alongside culture techniques (Fererra and Sanchez, 2016). Cultures provided the basis of our understanding of microorganisms, studies of such have provided the information (e.g. reference genomes and information from physiological studies) needed to understand and interpret data retrieved from ‘omics studies (Fererra and Sanches, 2016). With the introduction of single cell sorting and whole genome amplification (SAGs), culturing is no longer a necessity as this technique can be performed on non-pure culturable organisms.
‘Omics analysis techniques can be combined with experimentation to further improve our understanding of wastewater treatment processes.
References:
Ferrera, I., Sanchez, S. (2016). Insights into microbial diversity in wastewater treatment systems: How far have we come?. Biotechnology Advances, 34, 790-802. doi: 10.1016/j.biotechadv.2016.04.003
Stokholm-Bjerregaard, M., McIlroy, S. J., Mierychlo, M., Karst, S. M., Ablertsen, M., Nielsen, P. H. (2017). A Critical Assesment of the Microorganisms Proposed to be Important to Enhanced Biological Phosphorus Removal in Full-Scale Wastewater Treatment Systems. Frontiers in Microbiology, 8(718), 1-18. doi: 10.3389/fmicb.2017.00718